CloneAssist is designed to assist molecular biologists in expanding their workflow capabilities with sequence analyses, annotation, tracking, and reporting.
eLabNext is seeking users interested in becoming beta testers of this add-on to collect feedback for feature improvements. As beta testers, users will receive an extended 1-year free trial of CloneAssist. At its full release, CloneAssist will be offered as a paid add-on.
Become a beta tester
If you have any questions or are interested in becoming a beta tester, please contact your dedicated account manager or fill out our contact form.

CloneAssist Top Features – Overview
1). Seamlessly integrated with eLabNext
- The file is saved in a CloneAssist section, and a preview of CloneAssist is shown while the user is not working on it.
- Users may now create a CloneAssist section in an experiment.
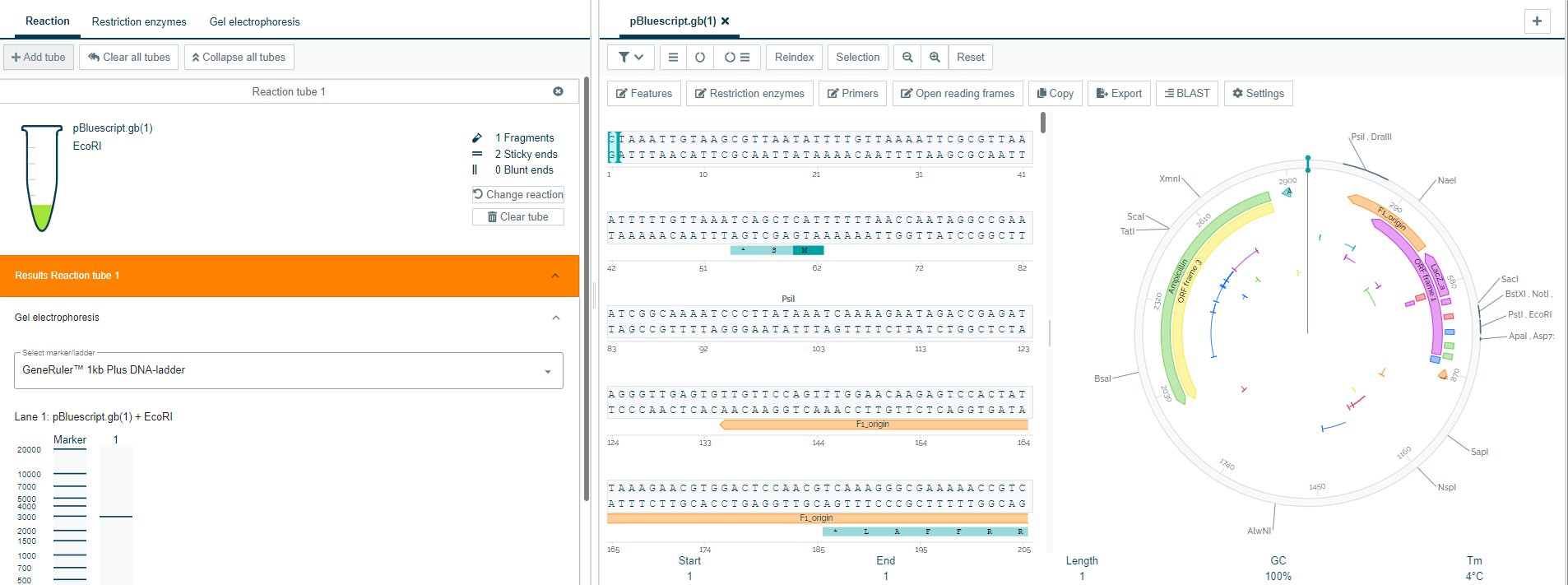
2). Plasmid visualization
- The data file is now visualized as a plasmid. This shows all features, primers, open reading frames, and restriction enzymes, making it easy to visualize the parts most relevant to your research.
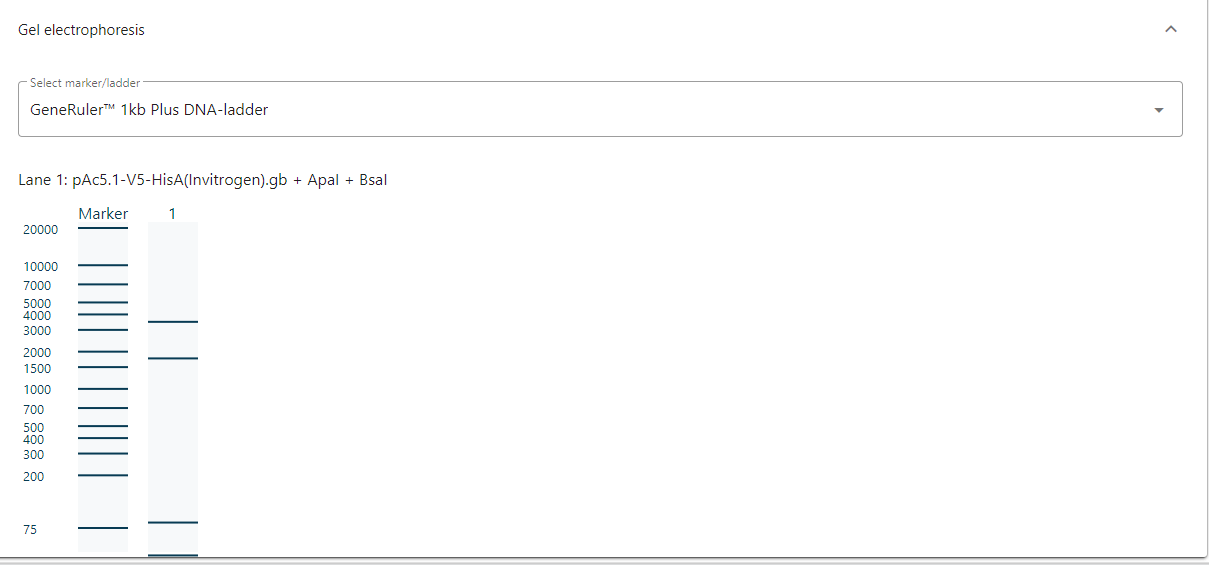
3). Gel electrophoresis
- By cutting your sequence with restriction enzymes, users get different fragments that are often verified with gel electrophoresis.
- CloneAssist provides a visualization of the gel electrophoresis with different markers.
- The gel can show one or more multiple reactions in one view.
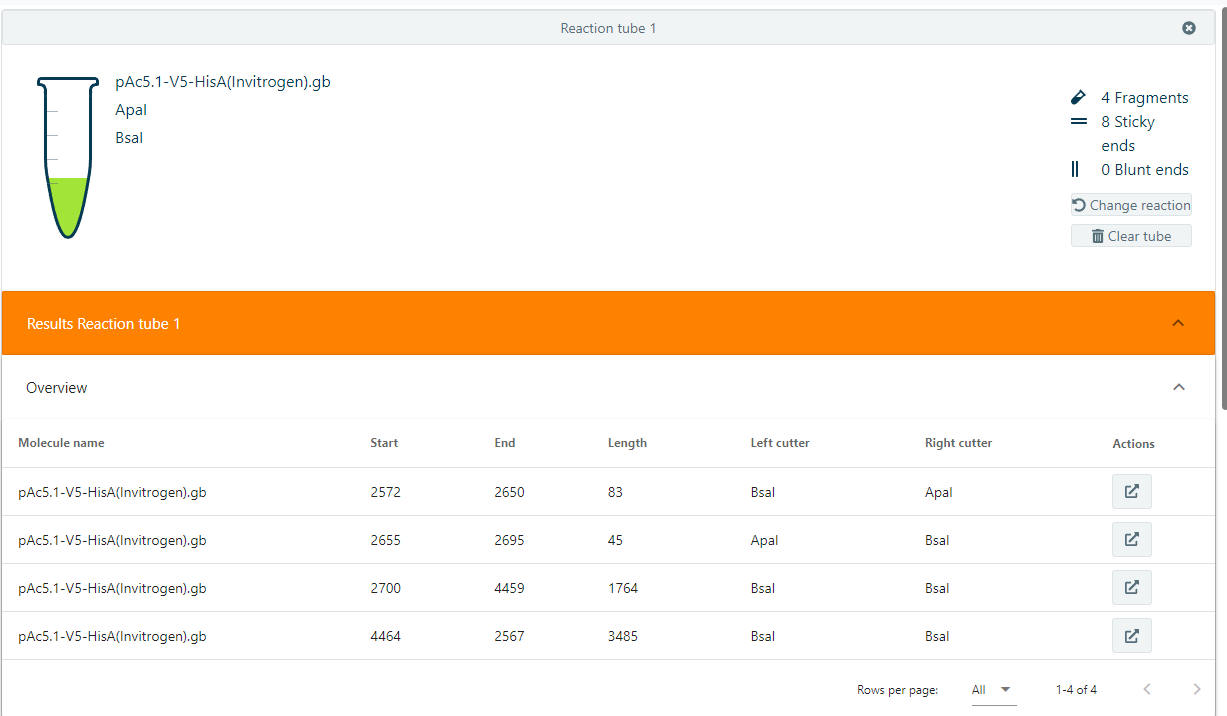
4). Easily cut molecules with certain restriction enzymes, then apply the results to new molecules
- When adding one or more restriction enzymes to your tube, users may directly see how many cuts the restriction enzyme will make in the molecule(s).
- Results show all fragments that have arisen from the cutting reaction.
5). Visualize both the Linear and Circular sequences
- When making a selection or selecting a feature of interest, the views are synchronized.
6). Add custom features
- Add your custom features in the sequence by hovering over the end of your selection and clicking ‘+ Feature’.
7). Work on multiple reactions at the same time
- Multiple tubes can be created to perform a reaction. They may be opened simultaneously or collapsed for future use.
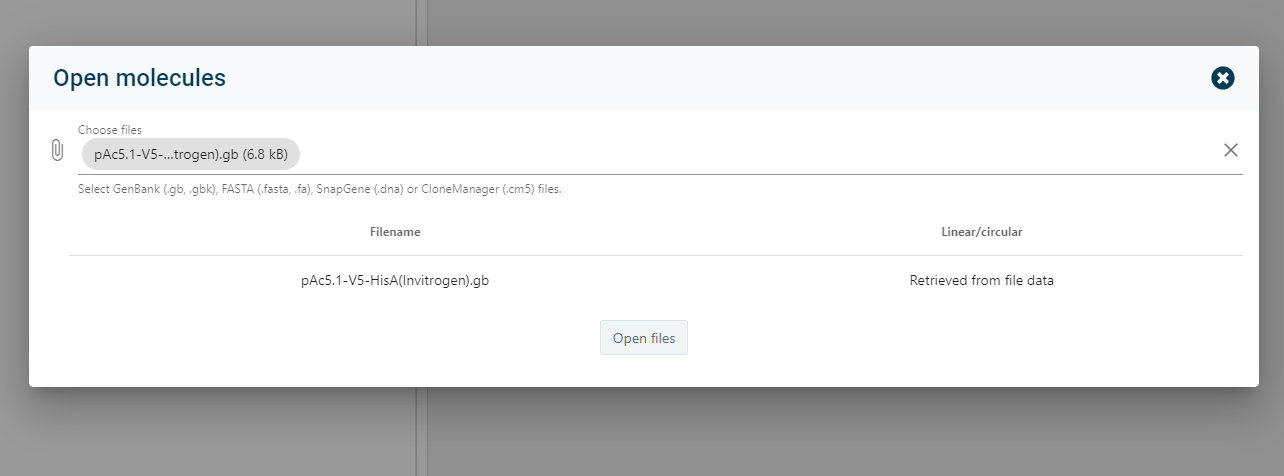
8). Multiple import/export options
- When starting a new CloneAssist section, you may import data from different formats such as snapgene (.dna), GenBank (.gb), CloneManager (.cm5), and fasta (.fasta).
- Molecules may be exported as an image (.png).
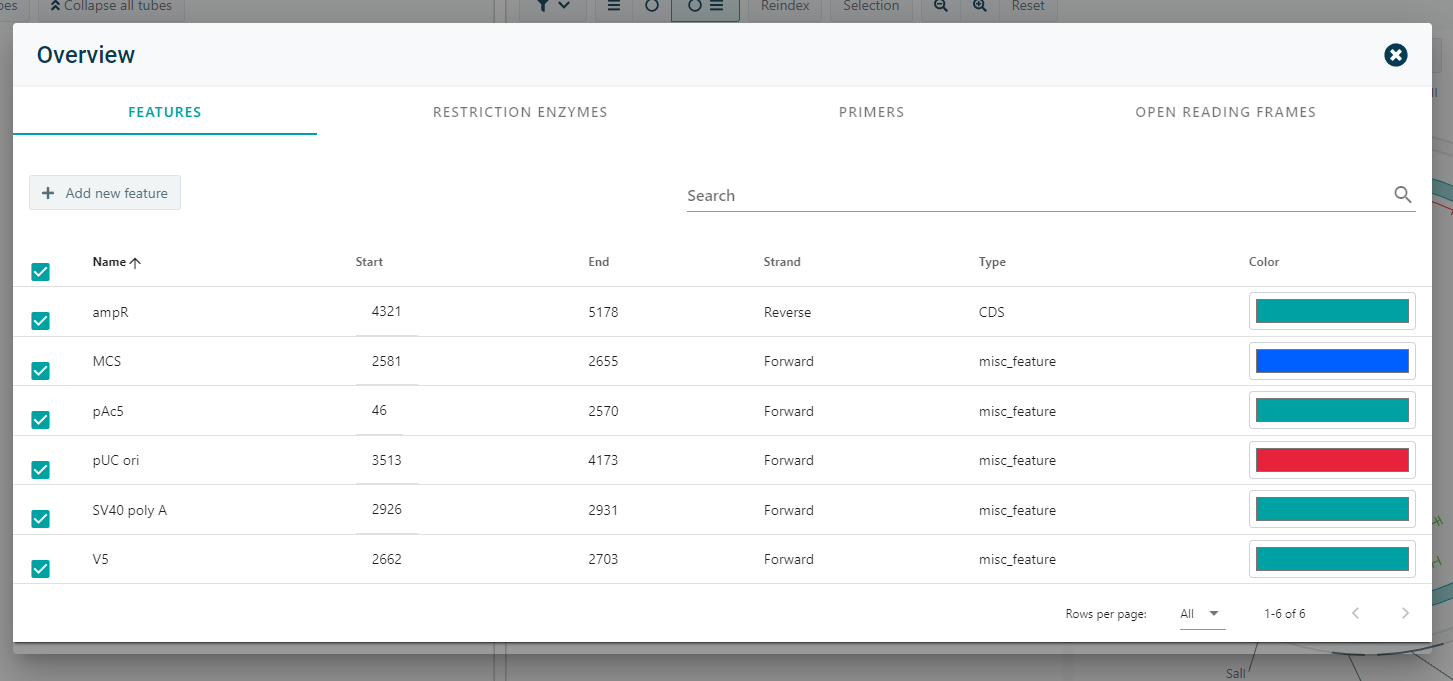
9). Filter the elements to the needs of your lab
- Visualizations are made to meet the needs of users. For example: if the user is only interested in the restriction enzymes that can cut the molecule, the user may disable all other visualization options.
- It is also possible to only show/hide specific features.
10). Sequence visualization shows, in great detail, how the DNA/RNA strand is constructed
- Linear View provides more detailed information, such as the actual sequence.
- When hovering over an enzyme, users can see the cutting sites.
These add-ons may also be useful

Signature Workflows
Enhance compliance and convenience by creating a custom approval workflows for signing and witness signing of experiments per project.
Learn more

Toxometris.ai
Accelerate your drug design process with cutting-edge in silico toxicity assessment
Learn more

AI Protocol Generator
AI Protocol Generator is your laboratory’s AI-powered protocol generation companion. Use this add-on to create complex protocols with just a…
Learn more